Flow cytometry is a laser-based technology used to analyze the physical and chemical characteristics of particles (typically cells) in a fluid as they pass through a laser beam. It enables multiparametric analysis of cell populations based on size, granularity, and fluorescence from labeled antibodies or dyes, at single-cell resolution and high throughput. A single-cell suspension is hydrodynamically focused and passed through a laser beam. Light scattering (forward scatter - FSC for size, side scatter - SSC for granularity) and fluorescence emissions from labeled cells are detected by photomultiplier tubes (PMTs), generating multi-dimensional data for each event.
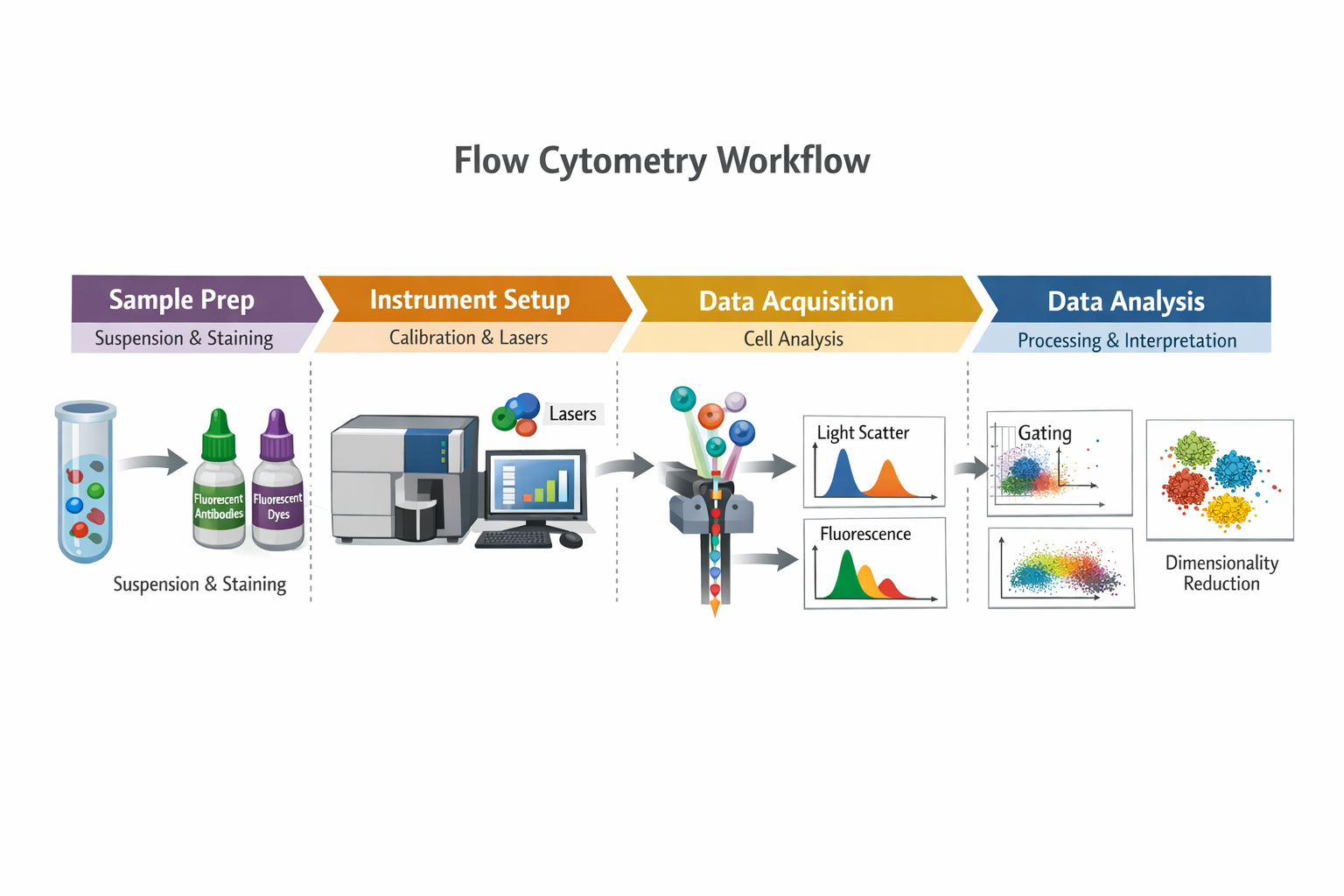
(Workflow Overview)
Stage 0: Experimental Design & Panel Setup
This critical planning stage occurs before any wet lab work.
1.1 Define Objectives
Identify target cell populations and markers (surface, intracellular, secreted).
1.2 Fluorophore Panel Design
1.2.1 Antigen Density.
Pair bright fluorophores (e.g., PE, APC) with low-expression markers; dim fluorophores (e.g., FITC, Alexa Fluor 488) with high-expression markers.
1.2.2 Spectral Overlap.
Use a spectrum viewer tool to minimize spillover.
1.2.3 Instrument Configuration.
Ensure your cytometer's lasers and filters can detect your chosen fluorophores.
1.2.4 Include Essential Controls.
See below.
Stage 1: Sample Preparation (Single-Cell Suspension)
Work on ice or at 4°C with pre-chilled buffers for live-cell staining..
1.1 Source Material.
1.1.1 Peripheral Blood / Bone Marrow.
Use RBC lysis buffer (e.g., ACK lysing buffer) or density gradient centrifugation (Ficoll-Paque) to isolate peripheral blood mononuclear cells (PBMCs). Wash 2x with FACS Buffer.
1.1.2 Cultured Cells (Adherent/Suspension).
Harvest using gentle enzymatic (e.g., Accutase) or non-enzymatic methods. Avoid trypsin if it cleaves your target epitopes.
1.1.3 Solid Tissues (Spleen, Lymph Node, Tumor).
Mechanically dissociate using a syringe plunger, gentleMACs dissociator, or manual grinding. Filter through a 40-70 µm cell strainer.
2.1 Wash & Count.
2.1.1 Centrifuge cell suspension at 300-500 x g for 5 minutes at 4°C. Decant supernatant.
2.1.2 Resuspend pellet in FACS Buffer (PBS + 1-2% FBS + 0.1% sodium azide (if not doing viability dye/functional assays)).
2.1.3 Count cells using a hemocytometer or automated counter.
2.1.4 Adjust concentration to 5-10 x 10⁶ cells/mL in FACS Buffer.
Stage 2: Cell Staining
Part A: Surface Antigen Staining.
2.1 Aliquot Cells.
Distribute 50-100 µL of cell suspension (0.5-1 x 10⁶ cells) per tube/well of a V-bottom or U-bottom plate.
2.2 Fc Receptor Blocking (Critical for primary cells like immune cells).
Incubate cells with purified anti-mouse CD16/32 (for mouse) or human Fc block for 10-15 minutes on ice.
2.3 Antibody Staining.
2.3.1 Add pre-titrated fluorescent antibodies directly to the cell suspension. Mix gently.
2.3.2 Incubate for 20-30 minutes in the dark on ice.
2.4 Wash.
2.4.1 Add 2 mL of FACS Buffer, centrifuge at 500 x g for 5 minutes at 4°C.
2.4.2 Aspirate supernatant completely. Resuspend pellet in 200 µL FACS Buffer for acquisition.
2.4.3 Proceed to acquisition immediately or fix cells (see below).
Part B: Intracellular / Intranuclear Staining (ICS/INS)
Requires prior surface staining, fixation, and permeabilization.
2.1 Fixation.
After surface staining and wash, resuspend cells in 100 µL of fixation buffer (e.g., 4% PFA, BD Cytofix). Incubate 20 minutes at RT or overnight at 4°C in the dark.
2.2 Wash.
Centrifuge, aspirate, and wash 1x with FACS Buffer.
2.3 Permeabilization.
2.3.1 Resuspend cells in 100-200 µL of permeabilization buffer (e.g., BD Perm/Wash, FoxP3 Buffer, or 0.1% Triton X-100). Incubate 15-30 minutes at RT or on ice.
2.3.2 Wash 1-2x with permeabilization buffer.
2.4 Intracellular Staining.
2.4.1 Resuspend cells in permeabilization buffer containing the pre-titrated intracellular antibody (e.g., anti-cytokine, anti-transcription factor).
2.4.2 Incubate for 30-60 minutes in the dark at RT or 4°C.
2.5 Final Wash & Resuspension.
2.5.1 Wash 2x with permeabilization buffer, then 1x with FACS Buffer.
2.5.2 Resuspend in 200-500 µL FACS Buffer for acquisition.
Part C: Viability Dye Staining
2.1 Add before surface staining.
Add a viability dye (e.g., Zombie UV, Fixable Viability Dye eFluor 506) diluted in PBS to the cell pellet. Incubate for 15-30 minutes at RT in the dark.
2.2 Wash.
Wash 2x with FACS Buffer before proceeding to surface staining.
Stage 3: Data Acquisition on the Flow Cytometer
3.1 Instrument Startup & QC.
3.1.1 Power on cytometer and computer. Run startup/primer.
3.1.2 Perform daily quality control (QC) using calibration beads (e.g., CS&T beads) to ensure laser delays, fluidics, and PMT voltages are standardized.
3.2 Setup Acquisition Template.
3.2.1 Create a plot for FSC-A vs. SSC-A to gate on cells.
3.2.2 Create a plot for FSC-A vs. FSC-H to gate singlets.
3.2.3 Create histograms or dot plots for each fluorescence channel.
3.3 Voltage Optimization & Compensation.
3.3.1 Run an unstained control and single-stained controls. Adjust PMT voltages so that the negative population is in the first decade of the log scale.
3.3.2 Use the single-stained controls to set up spectral compensation (manually or automatically) to correct for fluorophore spillover.
3.4 Acquisition.
3.4.1 Run your FMO (fluorescence-minus-one) controls and fully stained samples.
3.4.2 Acquire a sufficient number of events (e.g., 10,000-100,000 events within your population of interest).
3.4.3 Save data as .fcs files.
Stage 4: Data Analysis
4.1 Gating Strategy (Sequential, hierarchical gating)
4.1.1 Gate 1: Cells → Exclude debris on FSC-A vs. SSC-A.
4.1.2 Gate 2: Singlets → Select single cells on FSC-A vs. FSC-H (or SSC-A vs. SSC-H).
4.1.3 Gate 3: Live Cells → Exclude viability dye-positive (dead) cells.
4.1.4 Gate 4: Lineage/Subpopulation → Use specific markers (e.g., CD3+ for T cells).
4.1.5 Target Population → Analyze expression of your markers of interest.
4.2 Use FMO Controls
FMO controls define the boundary for "positive" vs. "negative" staining for each marker, especially for dim populations.
4.3 Quantification
Report results as % of parent population and/or Median Fluorescence Intensity (MFI).
Troubleshooting Guide
Control Type | Purpose | Example |
Low Cell Yield / Clogging | Tissue not fully dissociated, cell clumps. | Filter through a finer strainer (30-40 µm), use DNAse I during dissociation, increase pipetting. |
High Background (Poor Resolution) | Insufficient washing, antibody concentration too high, insufficient Fc block. | Increase wash volume/frequency, titrate antibodies, use Fc block for primary cells. |
Low Signal | Antigen downregulated, poor antibody binding, fixation/permeabilization too harsh. | Check sample freshness, titrate antibody, optimize fixation/permeabilization protocol. |
Poor Viability | Harsh processing, prolonged staining on ice. | Process samples quickly, keep cells cold, use a gentle viability dye. |
Compensation Issues | Wrong single-stain control, population shifted. | Use the same cell type for controls as your experiment, ensure controls are bright enough. |