Western blotting is a fundamental analytical technique used to detect specific proteins within a complex biological sample. It combines the resolving power of SDS-PAGE gel electrophoresis with the specificity of antibody-based immunodetection.
The entire procedure can be divided into five sequential stages:
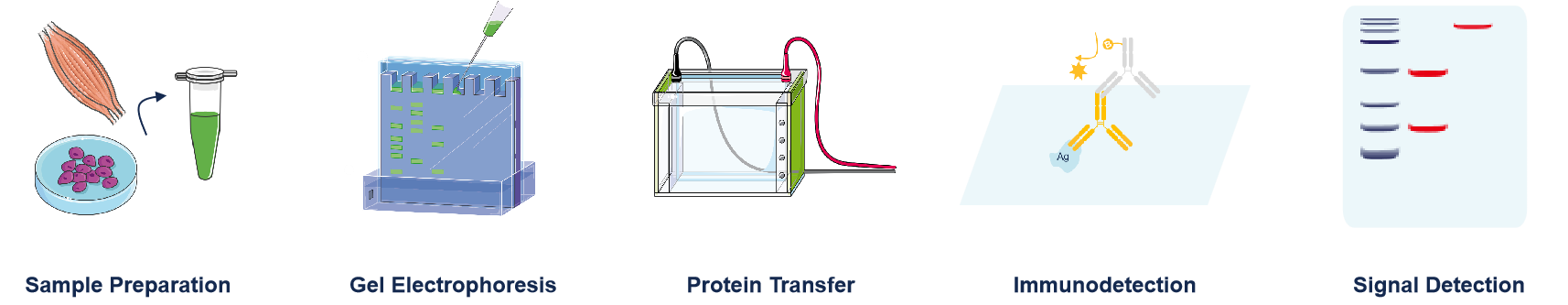
(Workflow Overview)
Stage 1: Sample Preparation & Protein Extraction
The goal is to obtain a clarified protein lysate where target proteins are soluble, intact, and represent their in vivo state.
1.1 Sample Collection
Adherent Cells: Wash with ice-cold PBS, then scrape or trypsinize into a collection tube on ice.
Suspension Cells: Pellet by centrifugation and wash with PBS.
Tissues: Snap-freeze in liquid nitrogen, then pulverize. Alternatively, homogenize directly in cold lysis buffer.
1.2 Cell/Tissue Lysis
Resuspend sample in appropriate, ice-cold lysis buffer.
Common Buffer Examples:
RIPA Buffer: General purpose, for total cell lysates.
Laemmli (SDS) Buffer: For direct denaturation and loading.
NP-40/Triton Buffer: Milder, for native protein complexes.
Essential Buffer Additives (add fresh):
Protease Inhibitor Cocktail
Phosphatase Inhibitor Cocktail (if studying phosphorylation)
PMSF (serine protease inhibitor)
1.3 Clarification
1.3.1 Incubate lysate on ice for 20-30 minutes.
1.3.2 Centrifuge at >12,000 x g for 15 minutes at 4°C.
1.3.3 Transfer supernatant (cleared lysate) to a new tube. Discard pellet.
1.4 Protein Quantification & Denaturation
1.4.1 Quantify: Measure protein concentration using BCA or Bradford assay.
1.4.2 Normalize: Dilute all samples to the same concentration.
1.4.3 Denature: Mix normalized lysate with 4X or 5X Laemmli Sample Buffer (containing SDS and DTT/β-mercaptoethanol).
1.4.4 Heat: Boil at 95-100°C for 5-10 minutes.
1.4.5 Samples can be used immediately or stored at -20°C/-80°C.
Stage 2: Gel Electrophoresis (SDS-PAGE)
Proteins are separated by molecular weight.
2.1 Gel Preparation
Cast a discontinuous polyacrylamide gel:
Resolving Gel (Lower): Higher % acrylamide (e.g., 8-15%). Separates proteins.
Stacking Gel (Upper): Lower % acrylamide (~4%). Concentrates proteins into a sharp line.
2.2 Loading & Running
2.2.1 Assemble gel in electrophoresis chamber filled with running buffer.
2.2.2 Load equal amounts (µg) of protein samples and a pre-stained protein ladder into wells.
2.2.3 Run at constant voltage (e.g., 80V through stacking gel, 120V through resolving gel) until dye front reaches bottom.
Stage 3: Protein Transfer (Blotting)
Proteins are moved from the gel onto a membrane.
3.1 Membrane & Buffer
Membrane Choice: PVDF (requires methanol activation) or Nitrocellulose.
Common Transfer Buffer: Tris-Glycine with 20% methanol.
3.2 Assembling the Transfer Stack
Construct the following sandwich from CATHODE (-) to ANODE (+):
Cathode Plate → Sponge → Filter Paper → Gel → Membrane → Filter Paper → Sponge → Anode Plate
Ensure no air bubbles between gel and membrane.
3.3 Transfer Methods
Wet/Tank Transfer: 100-400 mA, 60-90 min at 4°C. Robust for most proteins.
Semi-Dry Transfer: 15-25 V, 30-60 min. Faster, uses less buffer.
Stage 4: Immunodetection
Target protein is specifically identified using antibodies.
4.1 Blocking
Incubate membrane in blocking solution (e.g., 5% non-fat milk or BSA in TBST) for 1 hour at RT to prevent non-specific binding.
4.2 Primary Antibody Incubation
4.2.1 Dilute specific primary antibody in blocking buffer or antibody diluent.
Incubate membrane:
Option A: 1-2 hours at room temperature with agitation.
Option B (Recommended for sensitivity): Overnight at 4°C with gentle shaking.
4.2.2 Wash membrane 3 x 5-10 minutes with TBST.
4.3 Secondary Antibody Incubation
4.3.1 Incubate with HRP- or Fluorophore-conjugated secondary antibody (targeting the host species of the primary antibody) for 1 hour at RT.
4.3.2 Wash membrane thoroughly 3 x 5-10 minutes with TBST.
Stage 5: Signal Development & Analysis
5.1 Signal Detection
For HRP-conjugated Antibodies (Chemiluminescence):
Mix ECL substrate components.
Incubate membrane with substrate for 1-5 minutes.
Image using X-ray film or a digital CCD imager.
For Fluorescent Antibodies:
Rinse membrane.
Image directly using a fluorescence/scanner imager at appropriate wavelengths.
5.2 Membrane Stripping & Reprobing (Optional)
To detect a loading control (e.g., β-actin, GAPDH) on the same membrane, use a mild stripping buffer to remove antibodies, then re-block and re-probe.
5.3 Data Analysis
5.3.1 Estimate protein size using the molecular weight ladder.
5.3.2 Quantify band intensity using software (e.g., ImageJ, Image Studio Lite).
5.3.3 Normalize target protein signal to the loading control signal.
5.3.4 Compare relative expression levels between samples.